Over the last decade, Dr. Juan Zalapa has been working on a puzzle—a 500-million-piece one. Zalapa and others in his team—such as Luis Diaz-Garcia—is working to assemble the ‘Stevens’ cranberry genome in order to identify which genes control what traits in cranberry cultivars.
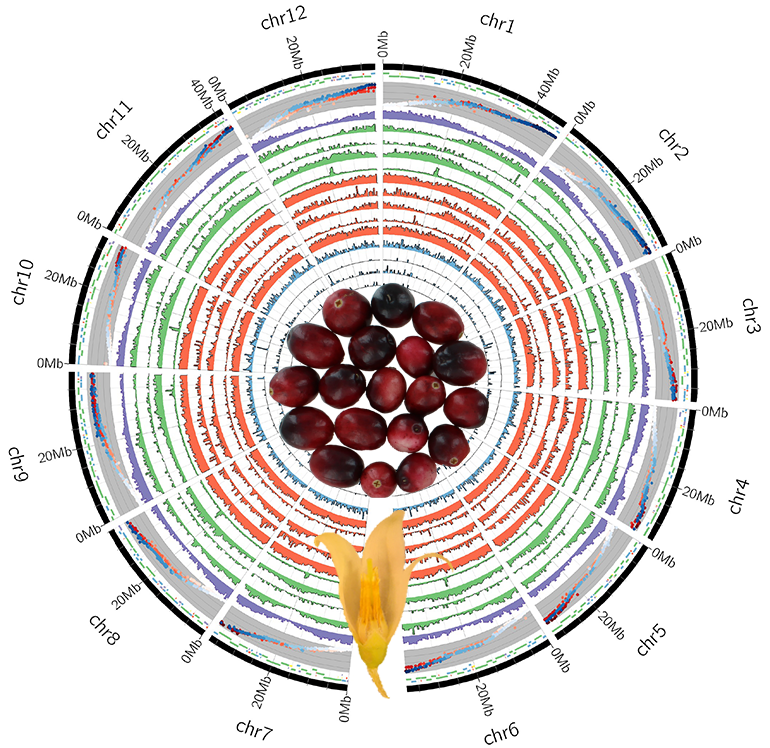
“Cranberries have 12 chromosomes. So, one of the things that we need to do for sequencing is to decipher the genetic code (bases) of those 12 chromosomes,” Zalapa said “When you do any kind of genetic sequencing, you can't get a single piece sequence for a whole chromosome. There are 500 million bases of chromosome sequence—divided into 12 chromosomes roughly, it could be 50 million bases per chromosome. And assembling this is like putting together a puzzle.”
Newer technology over the last ten years has made that puzzle assembly a bit easier and faster, allowing Zalapa and his team to obtain more and longer sequences at once and move closer to their sequence assembly goal. Zalapa and his team had to learn how to assemble or build the chromosomes back together, making improvements over the years before arriving at this current ‘Stevens’ genome iteration. And there is still some work to do for the sequencing project to refine and improve the chromosomes.
“There's still some gaps in different places of the chromosomes. And then we have some loose pieces here and there. So, this is still a chance for improvement,” Zalapa said. “That is why the VacCAP is going to continue to improve the assembly. And, in a couple years, we should have 10 more genomes of cultivars representing a wide diversity in cranberry. So, the Stevens cultivar genome is the first of many to come in.”
Having the ‘Stevens’ cranberry genome allows for more accurate marker selection in breeding the cranberries. While marker selection can be done with current linkage maps—roadmaps of general locations where certain genes are within the chromosomes—it isn’t as accurate.
“A molecular map is like a freeway or a highway, and you have signs that say there's a McDonald's coming up. But the sign is not the McDonald's, it is only announcing the exit to it is coming up,” Zalapa said. “So, the linkage map only tells you when something is around a general location. The cranberry genome actually gives me the exact location of something.”
Sequencing the genome is the key to finding the differences across cranberry cultivars that lead to different traits. According to Zalapa, there could be more than 99 percent similarity between cranberry cultivars, but that one percent or less that is different, is really what controls all of the differing traits for the cultivars—like high yield versus low yield, and high quality versus low quality.
“When we're talking about high fruit firmness, high yield, improved quality, or even the ability to survive under the adverse conditions or in a wider set of environments, all these traits are related to a very tiny amount of genetic information in the chromosomes, so we're talking about a sea of information that is the same, and only a few drops of water that matter,” Zalapa said. “And what [the ‘Stevens’ cranberry genome] allows us to do is match all of the things that are in common, and the things that are different among different cultivars. And then, we're going to start looking at what those differences are and what they actually do in terms of traits. This is why we need multiple diverse genomes sequenced, and this is the first of many genomes we have to do to understand the ‘good’ and ‘bad’ trait variation”
A better understanding of the cranberry genome and what genes control certain traits is important for an industry seeking to improve fruit quality. In time, this research will be invaluable for breeders and researchers to produce cultivars faster and more efficiently based on what traits the industry and the public demand.
“Through VacCAP, we're going to have multiple genomes; we will know more efficiently the chromosomes locations where the most important variation is that produce better cultivars and better traits” Zalapa said. “Once we know where the important chromosome locations are, we can start looking at the variation in traits in those areas. And this is going to accelerate the efficiency in selecting crosses and cultivars with the traits we want.”
The research was recently accepted for publication in Frontiers Plants Science journal as “Chromosome-level genome assembly of the American cranberry (Vaccinium macrocarpon Ait.) and its wild relative Vaccinium microcarpum”
The current data for the genome is also available on the Genome Database for Vaccinium (GDV) site for immediate use. This genome represents an extremely important resource for any ongoing and future cranberry genetic-genomic based projects or other projects seeking to effectively translate massive DNA and phenotypic datasets into useful information for the Vaccinium research and breeding community.













